library('available')
# help(package = 'available')
options(available.browse = FALSE)
available('merge', )
available('track')
available('trackChanges')
available('trackR')
available('collab')
available('writeTogether')Interesting R packages
R
Short notes about R packages
- Flow charts in R: ggflowchart
- Web scraping: rvest
- Spatial data / maps: Recommendations from Nicola Rennie’s blog
Check package names
R packages built from a single *.RMD file
Interfaces to computer algebra systems in R
Herre, I have collected a few resources in the field of computer algebra systems that have an interface to R.
General
- CRAN task view: Section “Multi-Precision Arithmetic and Symbolic Mathematics” on this site
- Blog article
Maxima
system('export PATH=${PATH}:/Applications/Maxima.app/Contents/Resources/opt/bin')
system('echo $PATH')
help(package='rim')library(rim)Maxima successfully registered as knitr engine!rim::maxima.isInstalled()[1] TRUErim::maxima.get("1+1;")(%o1) 2(%i1) lambda_S: beta * psi;(%o1) beta*psi(%i2) Lambda_S: integrate(lambda_S, t);(%o2) beta*psi*t(%i3) S_S: exp(-Lambda_S);(%o3) %e^-(beta*psi*t)(%i4) assume(beta > 0);(%o4) [beta > 0](%i5) assume(psi > 0);(%o5) [psi > 0](%i6) E_S: integrate(S_S, t, 0, inf);(%o6) 1/(beta*psi)(%i7) tex(''E_S);(%o7) falseRyacas
library(Ryacas)
Attaching package: 'Ryacas'The following object is masked from 'package:stats':
integrateThe following objects are masked from 'package:base':
%*%, det, diag, diag<-, lower.tri, upper.trias_r(yac_str("Simplify(a*b*a^2/b-a^3)"))[1] 0yac_str('Simplify(x^2 + 2*(x-3)^3)')[1] "2*x^3-17*x^2+54*x-54"yac_str("TeXForm(x^2 - 1)")[1] "x ^{2} - 1"Flowcharts
Source: https://cran.r-project.org/web/packages/Gmisc/vignettes/Grid-based_flowcharts.html
library(Gmisc, quietly = TRUE)
library(glue)
library(htmlTable)
library(grid)
library(magrittr)
org_cohort <- boxGrob(glue("Stockholm population",
"n = {pop}",
pop = txtInt(1632798),
.sep = "\n"))
eligible <- boxGrob(glue("Eligible",
"n = {pop}",
pop = txtInt(10032),
.sep = "\n"))
included <- boxGrob(glue("Randomized",
"n = {incl}",
incl = txtInt(122),
.sep = "\n"))
grp_a <- boxGrob(glue("Treatment A",
"n = {recr}",
recr = txtInt(43),
.sep = "\n"))
grp_b <- boxGrob(glue("Treatment B",
"n = {recr}",
recr = txtInt(122 - 43 - 30),
.sep = "\n"))
excluded <- boxGrob(glue("Excluded (n = {tot}):",
" - not interested: {uninterested}",
" - contra-indicated: {contra}",
tot = 30,
uninterested = 12,
contra = 30 - 12,
.sep = "\n"),
just = "left")
grid.newpage()
vert <- spreadVertical(org_cohort,
eligible = eligible,
included = included,
grps = grp_a)
grps <- alignVertical(reference = vert$grps,
grp_a, grp_b) %>%
spreadHorizontal()
vert$grps <- NULL
excluded <- moveBox(excluded,
x = .8,
y = coords(vert$included)$top + distance(vert$eligible, vert$included, half = TRUE, center = FALSE))
for (i in 1:(length(vert) - 1)) {
connectGrob(vert[[i]], vert[[i + 1]], type = "vert") %>%
print
}
connectGrob(vert$included, grps[[1]], type = "N")
connectGrob(vert$included, grps[[2]], type = "N")
connectGrob(vert$eligible, excluded, type = "L")
# Print boxes
vert
grps
excluded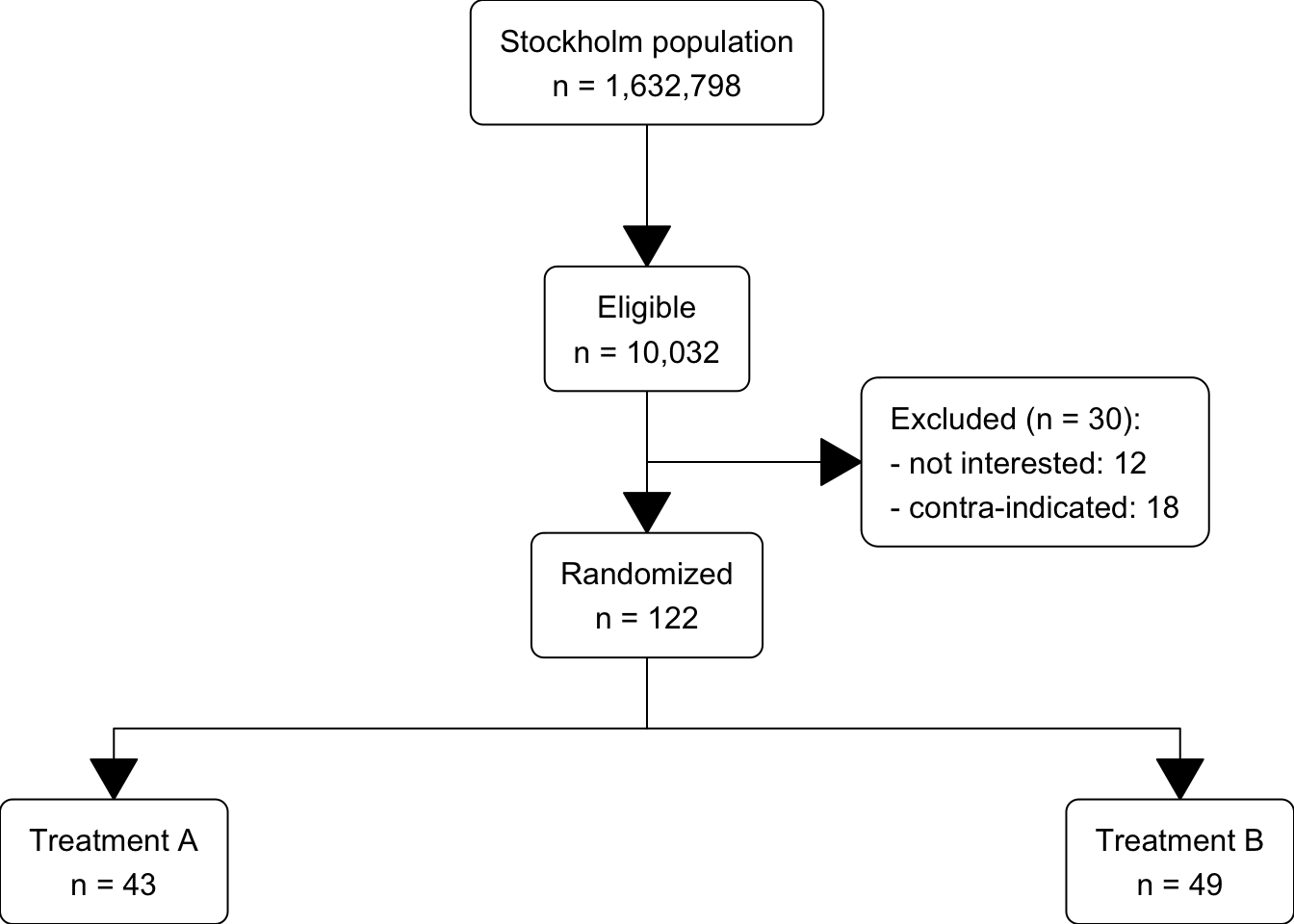
Leaflet for maps
Here for finding the path to the project directory
Package for formatting labels in ggplot
Concattenating text
Handle missing data
- Package nania - getting started
- However, there seem to be no methods for categorical data… → Opened issue
library('naniar')
library('tidyverse')── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.1.4 ✔ readr 2.1.4
✔ forcats 1.0.0 ✔ stringr 1.5.1
✔ ggplot2 3.5.1 ✔ tibble 3.2.1
✔ lubridate 1.9.3 ✔ tidyr 1.3.0
✔ purrr 1.0.2
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ tidyr::extract() masks magrittr::extract()
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
✖ purrr::set_names() masks magrittr::set_names()
✖ purrr::simplify() masks Ryacas::simplify()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorslibrary('simputation')
Attaching package: 'simputation'
The following object is masked from 'package:naniar':
impute_medianaq <- airquality |>
mutate(Ozone = as.double(Ozone),
fct = sample(c('a', 'b', NA), replace = TRUE, size = nrow(airquality)))
colSums(is.na(aq)) Ozone Solar.R Wind Temp Month Day fct
37 7 0 0 0 0 58 vis_miss(aq)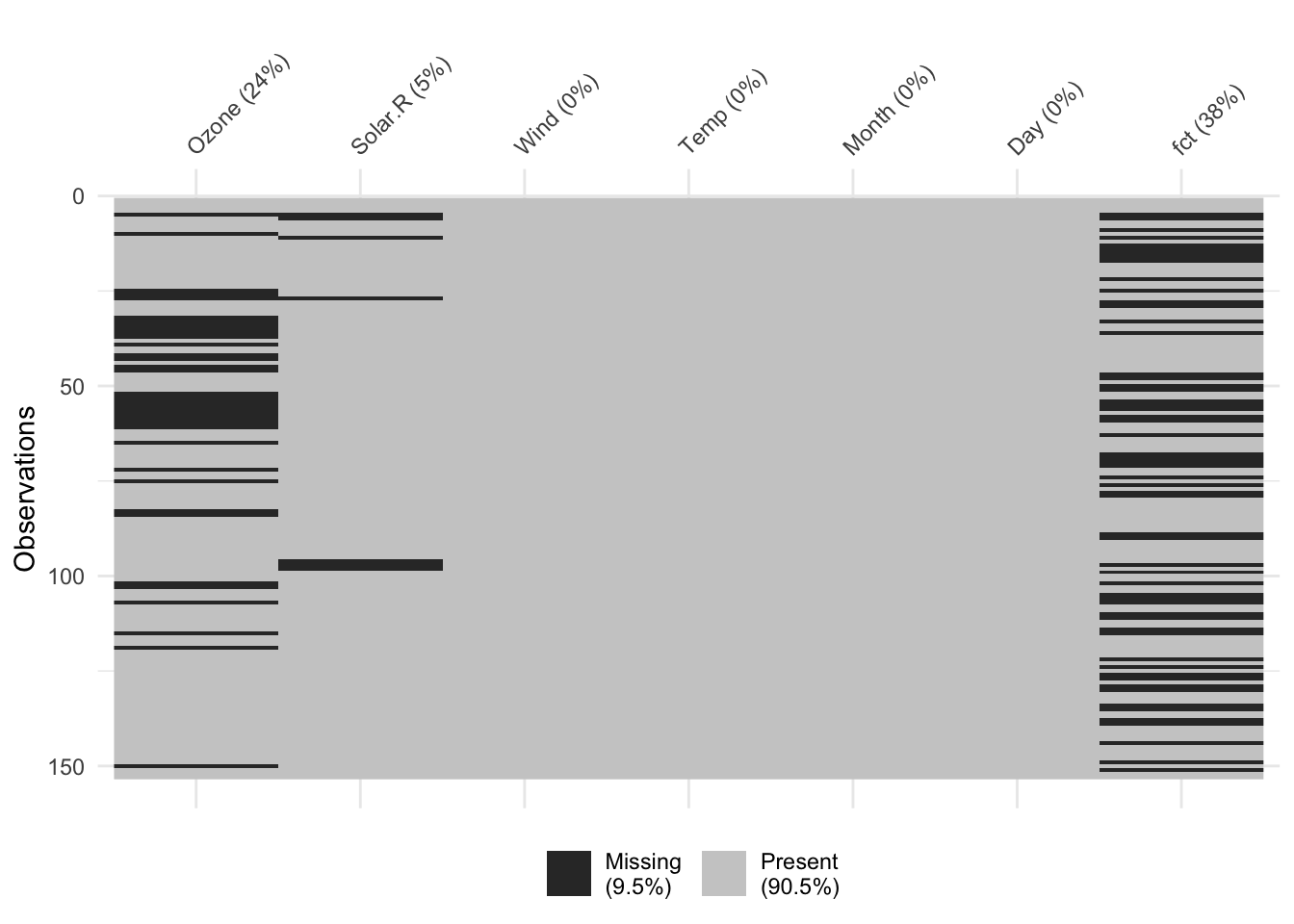
aq_nabular <- nabular(aq)
aq_nabular# A tibble: 153 × 14
Ozone Solar.R Wind Temp Month Day fct Ozone_NA Solar.R_NA Wind_NA
<dbl> <int> <dbl> <int> <int> <int> <chr> <fct> <fct> <fct>
1 41 190 7.4 67 5 1 a !NA !NA !NA
2 36 118 8 72 5 2 b !NA !NA !NA
3 12 149 12.6 74 5 3 b !NA !NA !NA
4 18 313 11.5 62 5 4 b !NA !NA !NA
5 NA NA 14.3 56 5 5 <NA> NA NA !NA
6 28 NA 14.9 66 5 6 <NA> !NA NA !NA
7 23 299 8.6 65 5 7 b !NA !NA !NA
8 19 99 13.8 59 5 8 a !NA !NA !NA
9 8 19 20.1 61 5 9 <NA> !NA !NA !NA
10 NA 194 8.6 69 5 10 a NA !NA !NA
# ℹ 143 more rows
# ℹ 4 more variables: Temp_NA <fct>, Month_NA <fct>, Day_NA <fct>, fct_NA <fct>aq_nabular |> impute_lm(Ozone ~ Temp)# A tibble: 153 × 14
Ozone Solar.R Wind Temp Month Day fct Ozone_NA Solar.R_NA Wind_NA
* <dbl> <int> <dbl> <int> <int> <int> <chr> <fct> <fct> <fct>
1 41 190 7.4 67 5 1 a !NA !NA !NA
2 36 118 8 72 5 2 b !NA !NA !NA
3 12 149 12.6 74 5 3 b !NA !NA !NA
4 18 313 11.5 62 5 4 b !NA !NA !NA
5 -11.0 NA 14.3 56 5 5 <NA> NA NA !NA
6 28 NA 14.9 66 5 6 <NA> !NA NA !NA
7 23 299 8.6 65 5 7 b !NA !NA !NA
8 19 99 13.8 59 5 8 a !NA !NA !NA
9 8 19 20.1 61 5 9 <NA> !NA !NA !NA
10 20.6 194 8.6 69 5 10 a NA !NA !NA
# ℹ 143 more rows
# ℹ 4 more variables: Temp_NA <fct>, Month_NA <fct>, Day_NA <fct>, fct_NA <fct>Collaborative writing
library("trackdown")Journal templates in quarto
- Link
- Can we add a template for Frontiers journals?
Debugging
library('icecream')
x <- 10
y <- 1:1000
ic(x + y)ℹ ic| `x + y`: num [1:1000] 11 12 13 14 15 16 17 18 19 20 ...ic(cars)ℹ ic| `cars`: data.frame [50 x 2]: $'speed': dbl [50], ...options(icecream.max.lines = 5,
icecream.prefix = "dbg:",
icecream.always.include.context = TRUE)
ic(x + y)ℹ dbg: <env: global> | `x + y`: num [1:1000] 11 12 13 14 15 16 17 18 19 20 ...options(icecream.peeking.function = utils::head,
icecream.max.lines = 5,
icecream.prefix = "dbg:",
icecream.always.include.context = TRUE)
ic(x + y)ℹ dbg: <env: global> | `x + y`: [1] 11 12 13 14 15 16ic(cars)ℹ dbg: <env: global> | `cars`:
speed dist
1 4 2
2 4 10
3 7 4
4 7 22Update R
Installed packages
dd <- installed.packages()
dd[, 'Package'] abind adaptr airports
"abind" "adaptr" "airports"
akima alphavantager anytime
"akima" "alphavantager" "anytime"
archive asd askpass
"archive" "asd" "askpass"
assertthat available babynames
"assertthat" "available" "babynames"
backports base base64enc
"backports" "base" "base64enc"
BayesFactor bayesplot bayestestR
"BayesFactor" "bayesplot" "bayestestR"
beanplot beeswarm betareg
"beanplot" "beeswarm" "betareg"
BH bigD bit
"BH" "bigD" "bit"
bit64 bitops blastula
"bit64" "bitops" "blastula"
blob bmp boot
"blob" "bmp" "boot"
box brew bridgesampling
"box" "brew" "bridgesampling"
brio brms Brobdingnag
"brio" "brms" "Brobdingnag"
broom broom.helpers bslib
"broom" "broom.helpers" "bslib"
cachem callr car
"cachem" "callr" "car"
carData caret caTools
"carData" "caret" "caTools"
catstats cellranger checkmate
"catstats" "cellranger" "checkmate"
cherryblossom circlize class
"cherryblossom" "circlize" "class"
classInt cli clinicaltrialr
"classInt" "cli" "clinicaltrialr"
clipr clisymbols clock
"clipr" "clisymbols" "clock"
cluster cmprsk coda
"cluster" "cmprsk" "coda"
codetools coin collections
"codetools" "coin" "collections"
colorspace colourpicker commonmark
"colorspace" "colourpicker" "commonmark"
compiler confintr conflicted
"compiler" "confintr" "conflicted"
contfrac coro corrplot
"contfrac" "coro" "corrplot"
COVID19 cowplot coxed
"COVID19" "cowplot" "coxed"
cpp11 crayon credentials
"cpp11" "crayon" "credentials"
crosstable crosstalk crul
"crosstable" "crosstalk" "crul"
cthist curl cyclocomp
"cthist" "curl" "cyclocomp"
DALEX DALEXtra data.table
"DALEX" "DALEXtra" "data.table"
datasets datawizard DBI
"datasets" "datawizard" "DBI"
dbplyr deldir DEoptimR
"dbplyr" "deldir" "DEoptimR"
Deriv desc DescTools
"Deriv" "desc" "DescTools"
deSolve devtools diagram
"deSolve" "devtools" "diagram"
dials DiceDesign diffobj
"dials" "DiceDesign" "diffobj"
diffviewer digest digitize
"diffviewer" "digest" "digitize"
distributional doParallel doRNG
"distributional" "doParallel" "doRNG"
dotCall64 downlit dplyr
"dotCall64" "downlit" "dplyr"
dqrng DT dtplyr
"dqrng" "DT" "dtplyr"
dygraphs e1071 effects
"dygraphs" "e1071" "effects"
effectsize effsize ellipsis
"effectsize" "effsize" "ellipsis"
elliptic emmeans estimability
"elliptic" "emmeans" "estimability"
evaluate Exact exactRankTests
"evaluate" "Exact" "exactRankTests"
expm fansi faraway
"expm" "fansi" "faraway"
farver fastai fastDummies
"farver" "fastai" "fastDummies"
fastmap fields filelock
"fastmap" "fields" "filelock"
fitdistrplus flexmix flextable
"fitdistrplus" "flexmix" "flextable"
FNN fontawesome fontBitstreamVera
"FNN" "fontawesome" "fontBitstreamVera"
fontLiberation fontquiver forcats
"fontLiberation" "fontquiver" "forcats"
foreach forecast foreign
"foreach" "forecast" "foreign"
forestplot Formula fracdiff
"forestplot" "Formula" "fracdiff"
fs furrr future
"fs" "furrr" "future"
future.apply gargle gdtools
"future.apply" "gargle" "gdtools"
generics geomtextpath gert
"generics" "geomtextpath" "gert"
getPass gfonts GGally
"getPass" "gfonts" "GGally"
ggbeeswarm ggdark ggdist
"ggbeeswarm" "ggdark" "ggdist"
ggeffects ggenealogy ggforce
"ggeffects" "ggenealogy" "ggforce"
ggformula ggmosaic ggpattern
"ggformula" "ggmosaic" "ggpattern"
ggplot2 ggpmisc ggpp
"ggplot2" "ggpmisc" "ggpp"
ggpubr ggquickeda ggraph
"ggpubr" "ggquickeda" "ggraph"
ggrepel ggridges ggsci
"ggrepel" "ggridges" "ggsci"
ggsignif ggstance ggstats
"ggsignif" "ggstance" "ggstats"
ggsurvfit ggtext ggthemes
"ggsurvfit" "ggtext" "ggthemes"
ggVennDiagram gh GillespieSSA
"ggVennDiagram" "gh" "GillespieSSA"
gitcreds gld glmmTMB
"gitcreds" "gld" "glmmTMB"
glmnet GlobalOptions globals
"glmnet" "GlobalOptions" "globals"
glue Gmisc goftest
"glue" "Gmisc" "goftest"
googledrive googlesheets4 gower
"googledrive" "googlesheets4" "gower"
GPfit gplots graphics
"GPfit" "gplots" "graphics"
graphlayouts grDevices grid
"graphlayouts" "grDevices" "grid"
gridExtra gridpattern gridtext
"gridExtra" "gridpattern" "gridtext"
gt gtable gtools
"gt" "gtable" "gtools"
gtsummary hablar hardhat
"gtsummary" "hablar" "hardhat"
haven here hfhub
"haven" "here" "hfhub"
highr Hmisc hms
"highr" "Hmisc" "hms"
htmlTable htmltools htmlwidgets
"htmlTable" "htmltools" "htmlwidgets"
httpcode httpgd httpuv
"httpcode" "httpgd" "httpuv"
httr httr2 hypergeo
"httr" "httr2" "hypergeo"
iBreakDown ica icecream
"iBreakDown" "ica" "icecream"
ids igraph infer
"ids" "igraph" "infer"
ingredients ini inline
"ingredients" "ini" "inline"
insight ipred irlba
"insight" "ipred" "irlba"
isoband iterators itertools
"isoband" "iterators" "itertools"
janeaustenr janitor jmvcore
"janeaustenr" "janitor" "jmvcore"
jomo jpeg jquerylib
"jomo" "jpeg" "jquerylib"
jsonlite juicyjuice kableExtra
"jsonlite" "juicyjuice" "kableExtra"
KernSmooth kinship2 km.ci
"KernSmooth" "kinship2" "km.ci"
KMsurv knitr knitrProgressBar
"KMsurv" "knitr" "knitrProgressBar"
labeling labelled laeken
"labeling" "labelled" "laeken"
Lahman languageserver later
"Lahman" "languageserver" "later"
latex2exp lattice lava
"latex2exp" "lattice" "lava"
lazyeval learnr leiden
"lazyeval" "learnr" "leiden"
lgr lhs libcoin
"lgr" "lhs" "libcoin"
lifecycle lintr listenv
"lifecycle" "lintr" "listenv"
lme4 lmerTest lmodel2
"lme4" "lmerTest" "lmodel2"
lmom lmtest logger
"lmom" "lmtest" "logger"
loo lpSolve lubridate
"loo" "lpSolve" "lubridate"
magick magrittr maps
"magick" "magrittr" "maps"
markdown MASS mathjaxr
"markdown" "MASS" "mathjaxr"
Matrix MatrixModels matrixStats
"Matrix" "MatrixModels" "matrixStats"
maxstat Mediana mediation
"maxstat" "Mediana" "mediation"
memoise metadat metafor
"memoise" "metadat" "metafor"
methods mgcv mice
"methods" "mgcv" "mice"
microbenchmark mime minhub
"microbenchmark" "mime" "minhub"
miniUI minqa missForest
"miniUI" "minqa" "missForest"
mitml mitools mlbench
"mitml" "mitools" "mlbench"
mlr3 mlr3measures mlr3misc
"mlr3" "mlr3measures" "mlr3misc"
mnormt modeldata modelenv
"mnormt" "modeldata" "modelenv"
ModelMetrics modelr modeltools
"ModelMetrics" "modelr" "modeltools"
mosaic mosaicCore mosaicData
"mosaic" "mosaicCore" "mosaicData"
msm mstate multcomp
"msm" "mstate" "multcomp"
multcompView MuMIn munsell
"multcompView" "MuMIn" "munsell"
mvtnorm naniar neuralnet
"mvtnorm" "naniar" "neuralnet"
nleqslv nlme nloptr
"nleqslv" "nlme" "nloptr"
nnet norm numDeriv
"nnet" "norm" "numDeriv"
nycflights13 officer openintro
"nycflights13" "officer" "openintro"
openssl ordinal packrat
"openssl" "ordinal" "packrat"
padr palmerpenguins pan
"padr" "palmerpenguins" "pan"
paradox parallel parallelly
"paradox" "parallel" "parallelly"
parameters parsnip patchwork
"parameters" "parsnip" "patchwork"
pbapply pbkrtest performance
"pbapply" "pbkrtest" "performance"
PerformanceAnalytics PermAlgo permute
"PerformanceAnalytics" "PermAlgo" "permute"
pheatmap pillar pkgbuild
"pheatmap" "pillar" "pkgbuild"
pkgconfig pkgdown pkgload
"pkgconfig" "pkgdown" "pkgload"
plogr plotly plyr
"plogr" "plotly" "plyr"
png polspline polyclip
"png" "polspline" "polyclip"
polynom posterior praise
"polynom" "posterior" "praise"
prettyunits pROC processx
"prettyunits" "pROC" "processx"
prodlim productplots profvis
"prodlim" "productplots" "profvis"
progress progressr promises
"progress" "progressr" "promises"
proxy PRROC ps
"proxy" "PRROC" "ps"
psych psycho purrr
"psych" "psycho" "purrr"
quadprog Quandl quantmod
"quadprog" "Quandl" "quantmod"
quantreg quarto QuickJSR
"quantreg" "quarto" "QuickJSR"
R.cache R.methodsS3 R.oo
"R.cache" "R.methodsS3" "R.oo"
R.utils R6 ragg
"R.utils" "R6" "ragg"
randomForest ranger RANN
"randomForest" "ranger" "RANN"
rappdirs rcmdcheck RColorBrewer
"rappdirs" "rcmdcheck" "RColorBrewer"
Rcpp RcppAnnoy RcppArmadillo
"Rcpp" "RcppAnnoy" "RcppArmadillo"
RcppEigen RcppHNSW RcppParallel
"RcppEigen" "RcppHNSW" "RcppParallel"
RcppProgress RcppRoll RcppTOML
"RcppProgress" "RcppRoll" "RcppTOML"
reactable reactR readbitmap
"reactable" "reactR" "readbitmap"
readODS readr readxl
"readODS" "readr" "readxl"
recipes rematch rematch2
"recipes" "rematch" "rematch2"
remotes renv repr
"remotes" "renv" "repr"
reprex reshape reshape2
"reprex" "reshape" "reshape2"
ResourceSelection reticulate rex
"ResourceSelection" "reticulate" "rex"
rgl riingo rim
"rgl" "riingo" "rim"
rjags rjson rlang
"rjags" "rjson" "rlang"
rmarkdown rms rngtools
"rmarkdown" "rms" "rngtools"
robustbase ROCR rootSolve
"robustbase" "ROCR" "rootSolve"
roxygen2 rpart rpart.plot
"roxygen2" "rpart" "rpart.plot"
RPostgres RPostgreSQL rprojroot
"RPostgres" "RPostgreSQL" "rprojroot"
rsample rsconnect RSpectra
"rsample" "rsconnect" "RSpectra"
rstan rstantools rstatix
"rstan" "rstantools" "rstatix"
rstudioapi rticles Rtsne
"rstudioapi" "rticles" "Rtsne"
RVenn rversions rvest
"RVenn" "rversions" "rvest"
Ryacas s2 safetensors
"Ryacas" "s2" "safetensors"
sandwich sass scales
"sandwich" "sass" "scales"
scattermore scatterplot3d scholar
"scattermore" "scatterplot3d" "scholar"
scico sctransform selectr
"scico" "sctransform" "selectr"
sessioninfo Seurat SeuratObject
"sessioninfo" "Seurat" "SeuratObject"
sf shape shiny
"sf" "shape" "shiny"
shiny.fluent shiny.react shinydashboard
"shiny.fluent" "shiny.react" "shinydashboard"
shinyFiles shinyjqui shinyjs
"shinyFiles" "shinyjqui" "shinyjs"
shinylive shinystan shinysurveys
"shinylive" "shinystan" "shinysurveys"
shinythemes simputation sitmo
"shinythemes" "simputation" "sitmo"
sjlabelled sjmisc sjPlot
"sjlabelled" "sjmisc" "sjPlot"
sjstats skimr slider
"sjstats" "skimr" "slider"
smisc snakecase SnowballC
"smisc" "snakecase" "SnowballC"
sodium sourcetools sp
"sodium" "sourcetools" "sp"
spam SparseM spatial
"spam" "SparseM" "spatial"
spatstat.data spatstat.explore spatstat.geom
"spatstat.data" "spatstat.explore" "spatstat.geom"
spatstat.random spatstat.sparse spatstat.utils
"spatstat.random" "spatstat.sparse" "spatstat.utils"
splines splus2R SQUAREM
"splines" "splus2R" "SQUAREM"
sscn StanHeaders statmod
"sscn" "StanHeaders" "statmod"
stats stats4 stringdist
"stats" "stats4" "stringdist"
stringi stringr styler
"stringi" "stringr" "styler"
survey survival survminer
"survey" "survival" "survminer"
survMisc svglite sys
"survMisc" "svglite" "sys"
systemfonts table1 tcltk
"systemfonts" "table1" "tcltk"
tensor tensorA testthat
"tensor" "tensorA" "testthat"
textshaping TH.data threejs
"textshaping" "TH.data" "threejs"
tibble tictoc tidycmprsk
"tibble" "tictoc" "tidycmprsk"
tidygraph tidymodels tidyquant
"tidygraph" "tidymodels" "tidyquant"
tidyr tidyselect tidytext
"tidyr" "tidyselect" "tidytext"
tidyverse tiff timechange
"tidyverse" "tiff" "timechange"
timeDate timetk tinytex
"timeDate" "timetk" "tinytex"
TMB tok tokenizers
"TMB" "tok" "tokenizers"
tools torch TOSTER
"tools" "torch" "TOSTER"
trackdown triebeard tseries
"trackdown" "triebeard" "tseries"
tsfeatures TTR TumorImmuneModels
"tsfeatures" "TTR" "TumorImmuneModels"
tune tweenr tzdb
"tune" "tweenr" "tzdb"
ucminf units UpSetR
"ucminf" "units" "UpSetR"
urca urlchecker urltools
"urca" "urlchecker" "urltools"
usdata usethis utf8
"usdata" "usethis" "utf8"
utils uuid uwot
"utils" "uuid" "uwot"
V8 vcd vctrs
"V8" "vcd" "vctrs"
vegan VIM vipor
"vegan" "VIM" "vipor"
viridis viridisLite visdat
"viridis" "viridisLite" "visdat"
vroom waldo warp
"vroom" "waldo" "warp"
WDI wesanderson whisker
"WDI" "wesanderson" "whisker"
withr wk workflows
"withr" "wk" "workflows"
workflowsets writexl xfun
"workflowsets" "writexl" "xfun"
XML xml2 xmlparsedata
"XML" "xml2" "xmlparsedata"
xopen xtable xts
"xopen" "xtable" "xts"
yaml yardstick yarrr
"yaml" "yardstick" "yarrr"
yesno yulab.utils zeallot
"yesno" "yulab.utils" "zeallot"
zip zoib zoo
"zip" "zoib" "zoo" dput(dd[, 'Package'], file = paste0('installed_packages_', Sys.Date(), '.RData'))
dput(dd, file = paste0('installed_packages_with_version_', Sys.Date(), '.RData'))Interesting packages for annoting ggplot plots
Interesting packages for surveys
- shinysurveys
library('shinysurveys')
if (interactive()) demo_survey_multipage()
if (interactive()) demo_survey()